|
1/4/2023 0 Comments Sds page layers 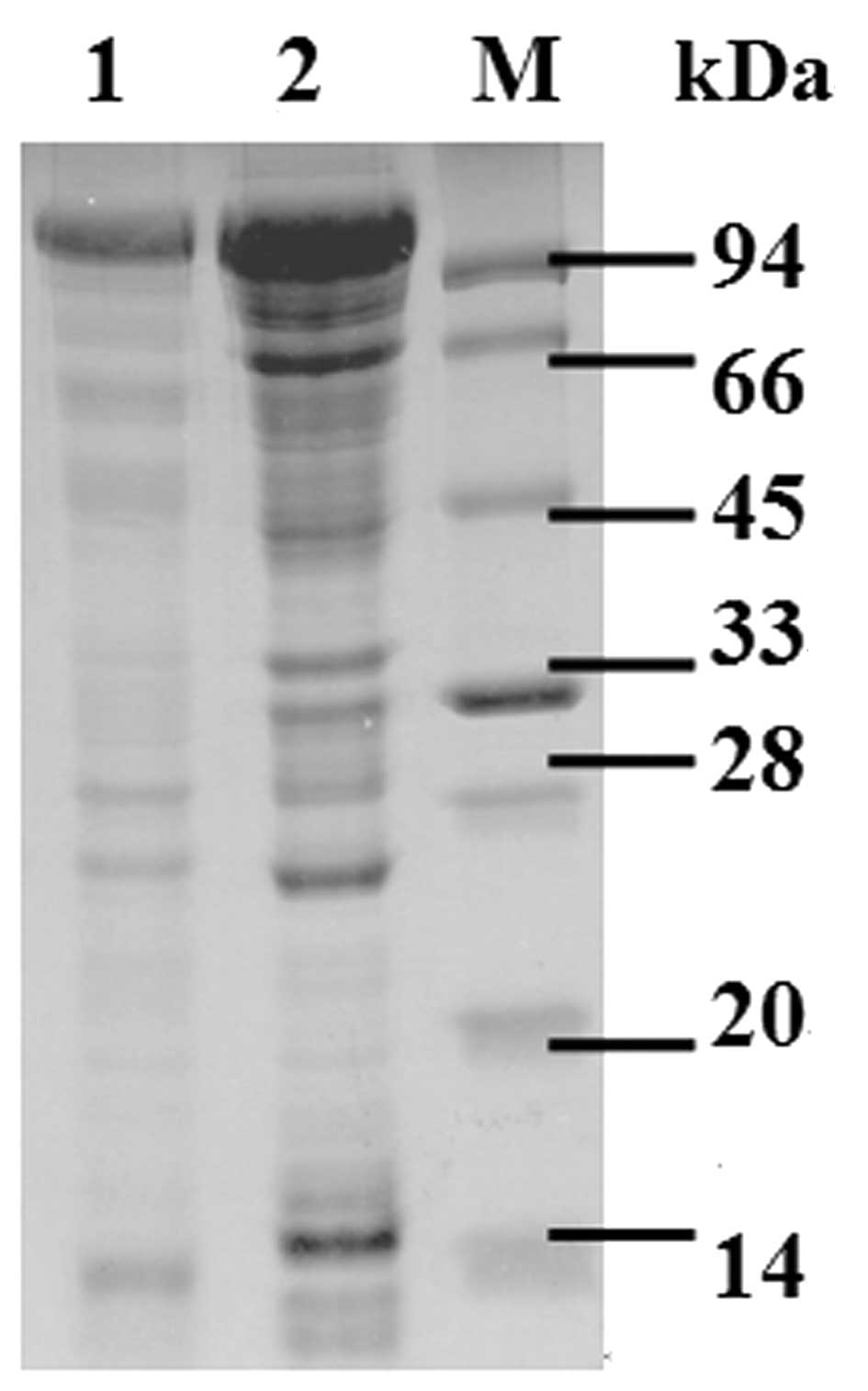
Schneider believes that this project has the potential to lead to groundbreaking changes in complete proteome mapping which will bring enormous benefits to the healthcare sector. As the protein translocates, the nanogap will be probed electrically and the amino acid sequence will be determined from the electrical readout of the graphene nanogap.ĭr. To sequence proteins, graphene will be sculpted into a nanogap (i.e., two layers of graphene separated by a physical gap on the order of the size of a protein molecule), through which unfolded proteins will be driven from head to tail under an electric field. Pour gel, leaving 2 cm below the bottom of the comb for the stacking gel. The stacking gel is a large pore polyacrylamide gel (4). There are two layers of gel, namely stacking or spacer gel, and resolving or separating gel. Graphene – a one atom thin material – has the potential to act as a sensor, primarily the edges of graphene. Mix in the following order: After adding TEMED and APS to the SDS-PAGE separation gel solution, the gel will polymerize quickly, so add these two reagents when ready to pour. Aims: In the present study, a method based on SDS-PAGE fingerprinting of surface layer proteins was developed to identify Lactobacillus delbrueckii subsp. SDS-PAGE SDS-PAGE, officially sodium dodecyl sulfate polyacrylamide gel electrophoresis, is a technique used in biochemistry, genetics and molecular. To date, sensing individual amino acids in proteins has been impossible because typical single-molecule nanopore sensors devices are thick (at least as thick as a protein). How SDS-PAGE works SDS-PAGE (sodium dodecyl sulphate-polyacrylamide gel electrophoresis) is commonly used in the lab for the separation of proteins based on their molecular weight. Grégory Schneider proposes to develop a new single molecule technique inspired from nanopores (widely used to study DNA), that can read the sequence of an individual protein from head-to- tail, amino acid per amino acid, molecule after molecule. Protein sequencing is hampered by three major roadblocks: i) proteins cannot be amplified as it is for DNA, ii) proteins have to be fragmented, and iii) unlike for DNA, 20 fluorescent amino acid tags without spectrum overlap simply does not exist. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
